Entrez is NCBI’s primary text search and retrieval system that integrates the literature and molecular databases at NCBI including DNA and protein sequence, structure, gene, genome, genetic variation, and gene expression. This document is an overview of the Entrez databases, with general information on searching and displaying data. More detailed help is available for the individual Entrez databases in the NCBI Help Manual sections on the NCBI bookshelf.
The Entrez search interface features powerful options for constructing precise searches and managing results. Options include popular configurable preset facet filters to help focus on specific kinds of results, an Advanced Search interface that facilitates constructing more sophisticated queries. Specialized search fields are available for each database and can be browsed and selected in the Search Builder section of the Advanced Search interface. Other useful Entrez features include Search History with access to recent results and a Clipboard where search results can be saved temporarily. A My NCBI account increases the power of the system by providing even more flexibility. Most importantly Entrez integrates data with links within and between databases. Not only does this interconnectivity enhance navigation and allow search results to be quickly focused or expanded; but also, more importantly, these relationships often expose unexpected connections that can lead to scientific discoveries.
The Entrez Databases
The Entrez system comprises over 30 different molecular and literature databases. New databases are added as biomedical science advances and new kinds of data become available. An alphabetical list of the current databases with a brief description of each is given below.
BioProject
The BioProject database is a searchable collection of complete and incomplete (in-progress) large-scale molecular projects including genome sequencing and assembly, transcriptome, metagenomic, annotation, expression, and mapping projects. BioProject provides a central point to link to all data associated with a project in the NCBI molecular and literature databases.
BioSample
BioSample contains descriptions of biological source materials used in studies that have data in other NCBI molecular databases such as Nucleotide and SRA.
Bookshelf
The NCBI Bookshelf contains a collection of full-text books that can be searched online and that are linked to PubMed records through research paper citations within the text. The collection includes biomedical textbooks, other scientific titles, and NCBI help manuals.
ClinVar
ClinVar is a public archive of submitted reports of clinically relevant human genetic variants and their relationships to phenotypes, with supporting evidence. ClinVar provides standardized nomenclature for variants and phenotypes, a review status for variants, and links to related NCBI literature and molecular databases.
Conserved Domains
Conserved Domains is a database of protein domains represented by sequence alignments and profiles for protein domains conserved in molecular evolution. It also includes alignments of the domains to known three-dimensional protein structures in the MMDB database. The source databases for Conserved Domains include Pfam, Smart, and COG.
dbGaP
dbGaP (Database of Genotypes and Phenotypes) provides the results of studies that have investigated the interaction of genotype and phenotype including genome-wide association studies, medical sequencing, molecular diagnostic assays, as well as association between genotype and non-clinical traits.
dbVAR
dbVAR (Database of Genomic Structural Variation) contains information about large-scale genomic variation, including large insertions, deletions, translocations, and inversions. dbVar also provides associations of defined variants with phenotype information.
Gene
Gene is a searchable database of genes, focusing on genomes that have been completely sequenced and that have an active research community to contribute gene-specific data. Information in Gene records includes nomenclature, chromosomal localization, gene products and their attributes (e.g., protein interactions), associated markers, phenotypes, interactions, and links to citations, sequences, variation details, maps, expression reports, homologs, protein domain content, and external databases.
GEO Datasets
GEO Datasets stores curated gene expression and molecular abundance data sets assembled by NCBI from the Gene Expression Omnibus (GEO) repository of microarray data.
GEO Profiles
GEO Profiles stores individual gene expression and molecular abundance profiles assembled from the Gene Expression Omnibus (GEO) repository of microarray data.
GTR
The Genetic Testing Registry (GTR) is a repository for voluntary submissions of genetic test information by providers. The scope includes the purpose of the test, methodology, validity, evidence of the usefulness of the test, and laboratory contacts and credentials. GTR includes information from and links to NCBI resources such as Gene, ClinVar and MedGen as well as many resources from outside the NIH.
MedGen
MedGen is a portal to information about human disorders and other phenotypes having a genetic component. MedGen aggregates the wide variety of terms used for disorders into a specific concept. Each concept may have associated clinical findings, causative genetic variants and the genes in which they occur, available clinical and research tests, molecular resources, professional guidelines, original and review literature, consumer resources, clinical trials, and links to other related NCBI molecular and literature databases as well as non-NCBI resources.
MeSH
MeSH (Medical Subject Headings) is the National Library of Medicine's controlled vocabulary and classification system (ontology) used for indexing articles in PubMed. MeSH terminology provides a consistent way to retrieve information that may use different terminology for the same concepts. Searches in the Entrez MeSH database provide synonymous MeSH terms that can provide more useful results in PubMed. The MeSH database records show subheadings access the MeSH browser showing related concepts and hierarchical relationships among MeSH terms.
NLM Catalog
The NLM Catalog contains records for books, journals, audiovisuals, computer software, electronic resources, and other materials in the National Library of Medicine (NLM) collections. The old Journals database was merged into the NLM Catalog database, and the information once retrieved via Journals, is provided by the NLM Catalog. This includes data such as journal title, MEDLINE abbreviation, NLM ID, ISO abbreviation, or ISSN.
Nucleotide
The Nucleotide database contains all the sequence data from GenBank, EMBL, and DDBJ, the members of the International Nucleotide Sequence Databases Collaboration (INSDC). Nucleotide also includes NCBI-curated Reference Sequences (RefSeqs), submitted assemblies and annotations from the Third Party Annotation (TPA) database, and nucleotide sequences extracted from structure records from the Protein Databank (PDB).
OMIM
The OMIM (Online Mendelian Inheritance in Man) database allows searches of OMIM articles about human genes, genetic disorders, and other inherited traits. OMIM articles provide links to associated literature references, sequence records, maps, and related databases. The NCBI service provides searching capabilities. OMIM records are hosted and served by the independent OMIM site (www.omim.org).
Protein
The Protein database contains amino acid sequences created from the translations of coding regions provided on nucleotide records in GenBank, EMBL, and DDBJ, the members of the International Nucleotide Sequence Databases Collaboration (INSDC) as well as those from coding regions on NCBI Reference Sequences and the Third Party Annotation (TPA) database records. Protein records are also imported from the outside protein-only data sources Protein Information Resource (PIR), UniProtKB/Swiss-Prot, Protein Research Foundation (PRF). Protein sequences are also extracted from structure records from the Protein Data Bank (PDB).
Protein Clusters
Protein Clusters is a collection of related protein sequences (clusters) consisting of Reference Sequence proteins that are encoded by complete prokaryotic genomes as well those encoded eukaryotic organelle plasmids and genomes. The database provides easy access to annotation information, publications, domains, structures, external links, and analysis tools.
PubChem BioAssay
PubChem BioAssay is a database that contains bioactivity screens of chemical substances described in PubChem Substance. It provides searchable descriptions of each bioassay, including descriptions of the conditions and readouts specific to that screening procedure.
PubChem Compound
The PubChem Compound database contains unique, validated chemical structures (small molecules) that can be searched using names, synonyms or keywords. The compound records may link to more than one PubChem Substance record if different depositors supplied the same structure. Structures in PubChem Compounds are pre-clustered and cross-referenced by identity and similarity groups. Additionally, calculated properties and descriptors are available for searching and filtering of chemical structures. Compound records are linked to related PubChem Substance Records, PubMed citations, protein 3D structures, and biological screening results that are available in PubChem BioAssay.
PubChem Substance
The PubChem Substance database contains information on chemical substances including mixtures electronically submitted to PubChem by depositors. This includes any chemical structure information submitted, as well as chemical names, comments, and links to the depositor's web site.
PubMed
PubMed is database of citations and abstracts for biomedical literature from MEDLINE and additional life science journals. Links are provided when full text versions of the articles are available through PubMed Central or other websites.
PubMed Central
PubMed Central (PMC) is the U.S. National Library of Medicine's digital archive of life sciences journal literature. PMC contains full-text manuscripts deposited by authors and articles provided by the publisher.
SNP
The SNP (Single Nucleotide Polymorphism) database is a central repository for single nucleotide polymorphisms, microsatellites, and small-scale insertions and deletions. Both submitted SNPs and NCBI-produced non-redundant reference records (RefSNPs) that cluster reports of the same polymorphism from different sources are available. SNP also contains population-specific frequency and genotype data, experimental conditions, molecular context, and mapping information for both neutral polymorphisms and clinical mutations.
SRA
The SRA (Sequence Read Archive) contains sequencing data from the next generation sequencing platforms. SRA accepts and presents data from all current next-generation sequencing platforms.
Structure
The Structure or Molecular Modeling Database (MMDB) contains experimental data from crystallographic and NMR structure determinations. The data for MMDB are obtained from the Protein Data Bank (PDB). Structure records link to bibliographic information, the sequence databases, and to the NCBI taxonomy. iCn3D, the web based NCBI 3D structure viewer, allows for easy interactive visualization of molecular structures from Entrez.
Taxonomy
The Taxonomy database contains the names and phylogenetic lineages of the organisms that have molecular data in the NCBI databases. New taxa are added to the Taxonomy database as data are deposited for them. The taxonomy records include links to all molecular data for the organism or group as well as links to outside classification resources. Taxonomy names provide the major controlled vocabulary for classifying molecular data across the Entrez system.
Access to the Entrez System
Nearly all search boxes that appear on the NCBI site access the Entrez system. The search box at the top of the NCBI homepage is a convenient place to begin Entrez searches. With the default All Databases selection, the results are presented on the Global Query page. This page lists the Entrez databases and the corresponding number of records found by the query in each database. The databases are organized into six broad categories on the results page: Literature, Health, Genomes, Genes, Proteins and Chemicals. Of course, the Global Query page itself can be used to search all database by entering a simple search term or phrase in the Search NCBI query box. Clicking on the number or the adjacent database name in Global Query retrieves the results in that database.
The search box on the NCBI homepage also has a pull-down list that allows selection of any of the individual databases. Alternatively, searches can be launched from the individual Entrez database pages. Many of the database homepages are linked directly to the NCBI homepage from the Popular Resource box in the upper right. It is also easy to access the database homepages directly using the simplified addresses that are formed by adding the database name to that of the NCBI homepage. For example, the address for the gene database homepage is simply www.ncbi.nlm.nih.gov/gene. Searches launched from the database homepage allow for more precise search strategies tailored to the database. These can be constructed using Boolean operators and combinations of one of more search field limits as described below.
Entrez Searching Options
Entrez queries can be single words, short phrases, sentences, database identifiers, gene symbols, or names. Often simple searches can result in overwhelming numbers of results or even no results at all. There are several built-in Entrez features that can help in creating more effective queries. These include Boolean operators, query translation, and fielded searching using any of the indexed fields available for the database. Any of these can be used in manually writing and editing queries but are also incorporated into various aspects of the interface so that precise results are available without the need to write complex query statements. These aspects of the interface include facets, and an Advanced Search page with a Search Builder and Search History that can be used to generate more sophisticated queries. More details on these features and some examples are given below.
Using Boolean Operators
Boolean operators provide a way of generating precise queries that produce well-defined sets of results. The Boolean operators used in Entrez and how they work are as follows.
AND: Finds documents that contain terms on both sides of the operator terms, the intersection of both searches.
OR: Finds documents that contain either term, the union of both searches.
NOT: Finds documents that contain the term on the left but not the term on the right of the operator, the subtraction of the right-hand search from the one on the left.
Entrez requires the Boolean operator AND to be entered in uppercase. This is not required in all databases for the other two operators, but it is simplest to enter all of them in uppercase:
promoters OR response elements NOT human AND mammals
Entrez processes all Boolean operators in a left-to-right sequence. Enclosing individual concepts in parentheses changes this priority. The terms inside the parentheses are processed first as a unit and then incorporated into the overall strategy. For example, in the following search statement, the union of response element and promoter results is generated first and then is intersected with the result of the g1p3 search.
g1p3 AND (response element OR promoter)
Default Boolean Combinations and Phrase Searching
Individual search terms separated by spaces are normally automatically combined as if they were joined by AND operators. The query tp53 mouse always gives the intersection of a search for mouse and a search for tp53. Each Entrez database also has an indexed phrase list. If a multi-word search matches a phrase, then only the phrase is used. For instance, the query protein kinase c is treated as a complete phrase rather than as the intersection of the three terms. The phrase indices and behavior may be different for different databases. In some cases, enclosing search terms in quotes can override the automatic intersection of terms and force a phrase search. The results for the phrase insulin dependent in most Entrez databases change based on whether the phrase is in quotes or not. Although phrase searching is useful, it should be used with caution because enclosing search terms in quotes restricts the documents retrieved to only those documents with exact matches to the text string within the quotes. Quoting a phrase may also prevent automatic term mapping of the individual terms to controlled vocabularies such as Medical Subject Headings or Organism (Taxonomy).
Indexed Fields, Query Translation, and Automatic Term Mapping
To facilitate searching, various indices are created for each Entrez database. These indices include information extracted from aspects of the record known as fields. Some of these fields contain essentially free text while other such as those for database identifiers (Accession, PMID), MeSH, and Organism are tightly controlled. Default searches in Entrez are All Fields searches. This usually results in the largest number of returned records but can produce unwanted results. For example, a search in any of the molecular databases with the term horse finds all records that contain this word in a variety of contexts, and many have nothing to do with that animal. If the goal is to find records specifically associated with that species, restricting the search to a particular field produces a more useful set of results. The available Fields and their indexed terms in any Entrez database can be explored on the Advanced Search page as part of the Search Builder. The Advanced Search interface, described in a separate section below, is linked beneath the search box of any page in an Entrez database as shown below for Nucleotide.
After a search there is also a Create alert option above the Search box that allows saving a search strategy in a My NCBI account. My NCBI provides the ability to schedule saved searches to be run automatically. The My NCBI help manual has more information on Saved searches and the other features of My NCBI mentioned in this document.
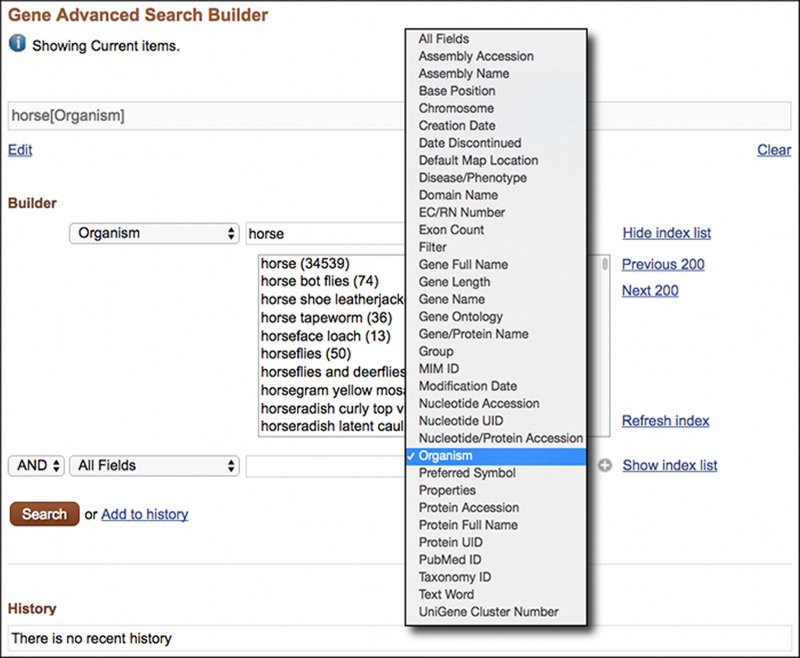
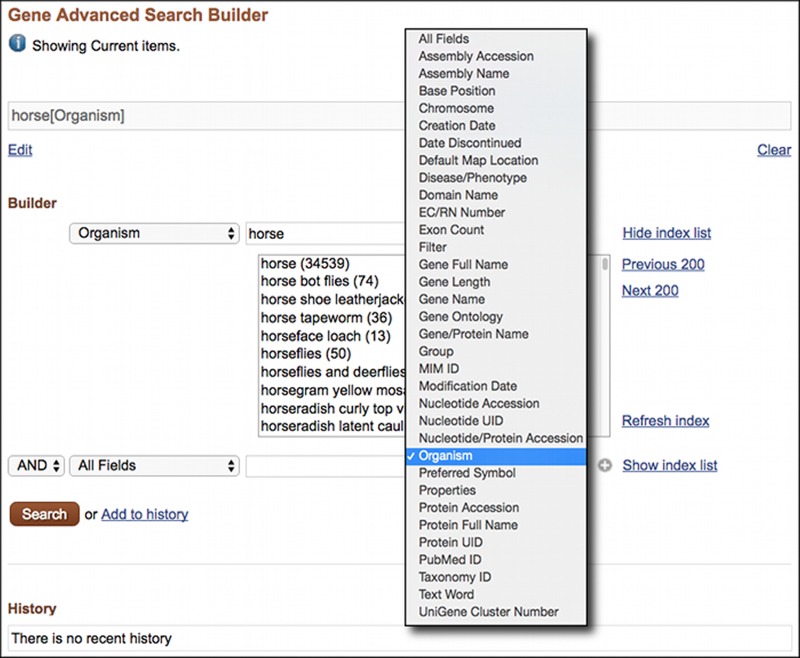
The Entrez Gene Advanced Search Page is shown in Figure 1 with the index for the Organism field expanded. The term horse is in the Organism index for Gene. Selecting the Organism field before the search finds only Gene records for the horse (Equus caballus) while the default All Fields search finds records for other species as well. Field restricted searching can be performed using the Search Builder. Restricted searches may also be entered manually by following the term with the name of the field in square brackets “[ ]”, as shown in the following examples:

Figure 1.
The Entrez Gene Advanced Search page showing the Search Builder with the Index for the Organism Field expanded.
horse[Organism]
neoplasms[MeSH Terms]
prolactin[Protein Name]
srcdb_refseq[Properties]
2010/06[Publication Date]
Dates and Other Ranges
Certain fields can accept ranges of values. Common examples are Publication Date, Modification Date, Accession, Molecular Weight, and Sequence Length. In these cases, the low and high numbers of the range are separated by a colon “:” as the range operator between them followed by the field:
110:500[Sequence Length]
2015/3/1:2016/4/30[Publication Date]
Facet Filters
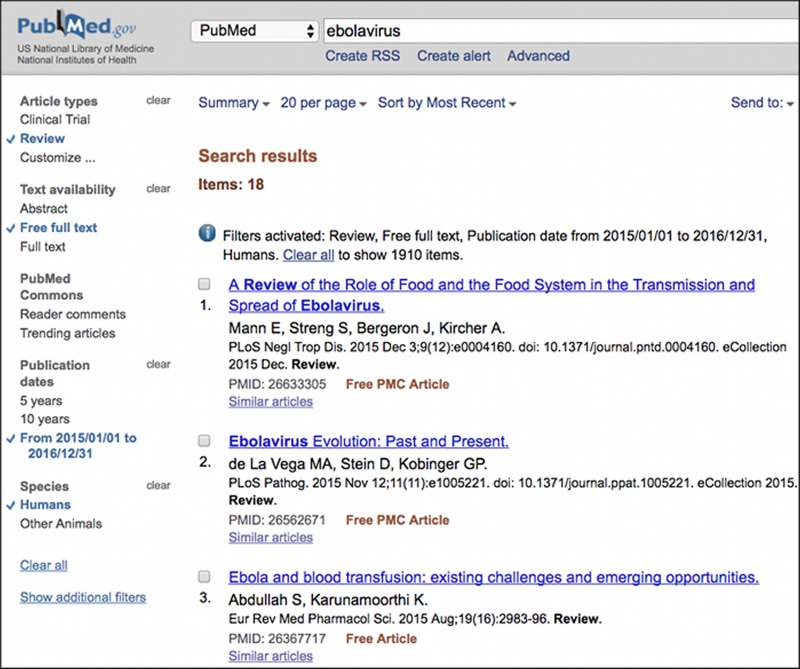
Many of useful restrictions including some of the search terms described above can be applied to searches through the faceted filter links on the left-hand column of the Entrez search page. Figure 2 shows a PubMed search page with several facets selected. Selecting any of the facet filters intersects the current search with the corresponding filter terms. Facet filters may restrict to certain types of records or exclude undesired ones.
Controlled Vocabulary Fields and Query Mapping
The indexed MeSH (Medical Subject Headings) and Organism fields have special roles in the PubMed and the traditional biomolecular databases respectively. Both MeSH and the Organism fields are tightly controlled vocabularies that are also hierarchical classification systems for the database records in PubMed and molecular databases. Every PubMed record is assigned sets of MeSH terms that add important information about the subject matter of the original paper. Records in the small molecule databases that are component PubChem are also associated with MeSH terms for the chemical components. The PubMed help document gives more details on the importance of the MeSH system. In a similar manner nearly all the biomolecular database records are attached to the source organism and its phylogenetic classification in the NCBI Taxonomy database. The MeSH Browser and the Taxonomy Browser are useful ways of exploring these systems and their associated records. Because of the importance of these two systems, queries are automatically mapped to these vocabularies whenever possible. The search terms may be expanded and translated as well. In certain databases other fields are involved in this mapping as well. The query horse dopamine receptor D2 becomes the more complex search statements in the PubMed and Protein Entrez search systems as shown in the box immediately below.
PubMed: ("horse s"[All Fields] OR "horses"[MeSH Terms] OR "horses"[All Fields] OR "horse"[All Fields]) AND ("receptors, dopamine"[MeSH Terms] OR ("receptors"[All Fields] AND "dopamine"[All Fields]) OR "dopamine receptors"[All Fields] OR ("dopamine"[All Fields] AND "receptor"[All Fields]) OR "dopamine receptor"[All Fields]) AND "D2"[All Fields]
Protein: ("Equus caballus"[Organism] OR horse[All Fields]) AND (dopamine
receptor D2[Protein Name] OR (dopamine[All Fields] AND receptor[All Fields]
AND D2[All Fields]))Special cases: Author names, Database Identifiers, and Stopwords
There are some other special cases of query interpretation in Entrez. Entering an author name in the form last name, first initials without punctuation, such as Lipman DJ, automatically maps to an author field search. Entering a recognized identifier for certain databases bypasses the general indices and directly retrieves the record. Identifiers that behave this way include accessions and gi numbers for sequence records, PubMed identifiers (PMIDs), and gene identifiers. Another special case is that certain words are ignored in Entrez searches. These words, known as stopwords, occur frequently in text on records but are not informative. Simple examples are definite and indefinite articles, conjunctions, and prepositions. A complete list of Entrez (PubMed) stopwords is given in the PubMed Guide. Punctuation in search terms is also typically ignored by Entrez and can cause certain strings to be missed. Enclosing problematic terms in quotes can help.
Using Wild Cards or Query Truncation
Entrez allows searching with single word stems where the ending of the term is replaced by an asterisk “*” to represent any character. This is often called truncation searching. For example, the search term hors* in the Protein database finds records with the terms hors, hors4, horse, horse's, horseradish, horst, and many more. Truncation is supported in fielded as well as All Fields searches and is helpful if the spelling of a word is uncertain. It can also help gather ranges of identifiers. For example, the following search statement in the Entrez Nucleotide database will find records for all human chromosomes:
NC_0000*[Accession] AND Human[Organism]
Because truncation searches use the only first 600 variations of a search term indexed for a particular field, poorly determined terms, for example cat* in PubMed, will give incomplete results.
Search Details Shows Query Interpretation
In some cases, it is useful to see how Entrez interpreted, expanded, or mapped the query as described above. In most databases, this information is provided in the Search Details in the Search details box in the right-hand column of search results. The Search details box below shows how the following search is interpreted in Protein.
Lee Y cytochrome oxidase 1 in the catfishes
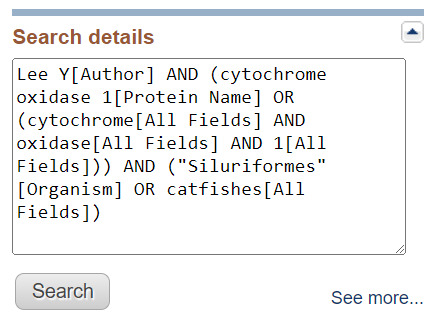
The words “in” and “the” are stopwords ignored by Entrez; “Lee Y” is mapped to an author search; and the term catfishes is expanded and mapped to the Organism vocabulary. The protein name “Cytochrome oxidase 1” is searched both as a phrase and as individual tokens.
Note: In PubMed the Search details are available on the Advanced Search page in the Search History.
Using the Advanced Search Page to Construct Complex Search Statements
The Advanced Search page for each Entrez database is useful for constructing complex and highly precise queries. Figure 3 shows the Advanced Search page for Entrez protein. The page functions as an independent search interface that allows formulation of complex queries. The Search Builder in combination with Search History facilitates the construction of more precise queries.
The pull-down list in the Search Builder shows all of the fields indexed for a particular database. The Show Index link opens an alphabetical list of terms for the selected field. When a term is entered in the Search Builder, the index will open to the closest match in the index. The Add to Search Box button puts the field-restricted queries into the Search Box. These may be run using the Search button or may be added to the Search History using the Preview button.
The Search History is maintained separately for each Entrez database and keeps track of all searches until the Web browser is closed or the history is deleted. Histories are automatically deleted after eight hours of inactivity. Entries in the Search History may be combined to create new searches that give precise results. The example in Figure 3 combines searches for frogs (#28), RefSeq proteins (#26), with a protein name search for prolactin to obtain the prolactin protein record for Xenopus tropicalis, NP_001093699.
Displaying and Saving a Set of Records
The Display Settings and Send to menus at the upper left and upper right of Entrez pages manage how records are displayed and stored or downloaded. The Display Settings menus have options for format, number of results per page, and sorting order. The available formats and sorting options vary depending on the database. The default format for multiple search results in Entrez is the Summary format that is consistent across databases. Single record default formats depend on the database. The default number of records displayed is 20 per page presented in the default sorting order for the database. These default settings may be modified by setting personal Preferences in a My NCBI account as described in the My NCBI Help book.
The Send to menu has options for sending results to online storage in Collections in My NCBI, the NCBI Clipboard for the database, or to a local file. Additional options may be available depending on the database. When choosing the file option, the record format and sorting order can be specified. By default, all Display Settings and Send to menu operations affect all records unless individual items are selected using the checkboxes at the left of the record title.
About the Clipboard and My Collections
The Clipboard is a temporary place on the NCBI website to save records. Each Entrez database has its own independent Clipboard that is limited to 500 items. Items saved to the clipboard are lost after eight hours of inactivity. When there are items in the clipboard, a link to access the clipboard appears in the upper right of any Entrez page for that database. The Clipboard behaves in the same way as any other page view in the database with equivalent Display Settings, Send to menus, and other features of that database. Records can be removed from the Clipboard through the link next to each item or by selecting the items using the check box and clicking the Remove Selected Items link at the top of the page. This link also functions to clear the Clipboard when no records are selected.
My Collections that is a part of the My NCBI service is a more permanent place to save records.
Related data: Neighbors and Links
One of the most useful and powerful features of the Entrez system is the integration of the data so that relationships between records can easily be explored. Many of these relationships may be unanticipated making the Entrez system an engine for scientific discovery. There are two major kinds of relationships established in the Entrez system: computationally derived associations within a database – items connected in this way are often called neighbors, and relationships based on information present on the records themselves, sometimes called hard links.
Neighbors are established automatically for some Entrez databases by computing on the data in the records. For example, related structures are determined using Vector Alignment Search Tool (VAST), a structure similarity algorithm; related PubMed citations (Similar articles) are identified by an algorithm that compares information rich words and phrases in the abstracts.
Reciprocal links between databases are established from the records themselves. For example, molecular records such as sequences or structures are linked to the PubMed citations where these data were reported. The literature citation has the reciprocal links back to the molecular databases. PubMed citations are linked to the full-text article in PubMed Central. Databases such as Gene that gather records from different sources have links back to their source records.
Combining neighbors and hard links can be an especially effective method for navigating across data and finding the most useful information. Moreover, this can be accomplished without performing additional searches. For example, going from an mRNA sequence in the nucleotide database to the three-dimensional structure for the corresponding or closely related protein can be a simple matter of following the link to protein, and follow the Related Structures link.
Access to Related Data
The Discovery Column
Easy access to related data is available on the individual records through the Links menu, the list of hypertext links available in the All links from this record section in the right-hand column of the full-record display. This right-hand column known as the Discovery Column contains additional access to related data and, in some cases, analysis tools. Many components in the Discovery column function as advertisements for related data providing more details about what information is available. The Discovery Column also functions in the search results providing additional options for searching and navigating including alternative search strategies as well as the Search details and Recent activity components described in the Entrez Searching section of this document. The Discovery Column on search results and other multiple record displays also provides access to related data through the Find related data component shown immediately below set for PubMed to Protein RefSeq. This follows the corresponding link for all records in the display when the Find items button is clicked.
In some databases, certain links to related data are also available from the individual summaries in search results, for example Identical Proteins in the Protein databases. Optionally, a preferences setting in My NCBI allows all links to be displayed in the summaries.
Recent Activity
An additional component that appears at the bottom of the Discovery Column is the Recent Activity list. While the Search History described previously in this document as part of the Advanced Search page, can be used to navigate to previous search results in the same database, it does not allow cross-database access. The Recent Activity component, present on many Entrez searches and record views, provides navigation to searches and record views in other databases. Only links to the last five searches and record views are listed by default, but activity for the past six months is available through a My NCBI account, where these searches can be saved and records can be added to collections.
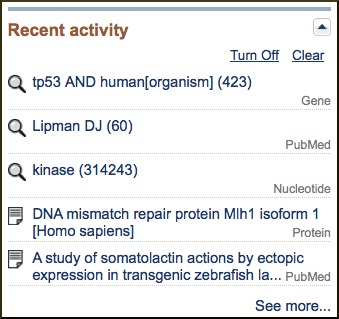
Creating Links to Web Pages in the Entrez System
The Entrez system now uses a standardized structure for web page addresses (URLs) that makes it easy to construct HTML links to address page displays and perform single searches. The standard URL format consists of the base URL for the database followed by options that can specify the record to be displayed, display options, and search terms. The Entrez Programming Utilities (E-Utilities) described in the next section of this document should be used to send frequent queries or retrieve large numbers of records from Entrez.
Database Homepages, Advanced Search, Limits
The base URL alone retrieves the homepage for the resource. The Advanced Search and Limits pages may also be addressed directly.
Examples:
Nucleotide homepage: www.ncbi.nlm.nih.gov/nucleotide
PubMed homepage: www.ncbi.nlm.nih.gov/pubmed
Gene Advanced Search: www.ncbi.nlm.nih.gov/gene/advanced
Linking to Records in the Entrez System
Linking Directly to Records Using an Identifier
The base URL for the database followed by a valid unique identifier for the database retrieves the record in the default single record view format. The summary format is displayed by default for more than one record. Most valid unique identifiers are purely numeric such as PubMed IDs, gene ids, and gi numbers for sequences. For the sequence databases accession numbers or accession.version numbers may also be used to address specific records. The report and format display options may be specified following the identifiers. The options available for report vary depending on the database and are listed on the Display settings menu for record views in that database. The NCBI Help Manual sections on individual Entrez databases have more information on available report formats. The format option may be used to retrieve text rather than the default HTML format.
Examples:
Protein gi 4557757, GenPept format (default)
www.ncbi.nlm.nih.gov/protein/4557757
Nucleotide accessions NM_000240 and NM_000041 in GenBank format
http://www.ncbi.nlm.nih.gov/nucleotide/NM_000240,NM_000041?report=genbank
Gene ID 348 full report (default)
Gene ID 348 in XML format
Linking to Search Results
Links to search results in Entrez databases may be created by adding a term to the URL. Report options and the maximum number of records to display per page may also be specified.
Examples:
Search in nucleotide for APOE with gene field restriction and 200 records displayed:
www.ncbi.nlm.nih.gov/nucleotide/?term=APOE[gene]&dispmax=200
Search in PubMed for Lipman DJ:
Programmatic Access to the Entrez System
The Entrez system may be accessed programmatically for high volume non-interactive search and retrieval through the Entrez Programming Utilities (E-utilities). These are a set of eight server-side programs that provide a stable interface to the Entrez query and database system. More information E-utilities is available in the Entrez Programming Utilities Help manual on the NCBI Bookshelf.
Publication Details
Author Information and Affiliations
Authors
Peter Cooper, PhD1 and Monica Romiti, MLIS2.Affiliations
Publication History
Created: January 20, 2006; Last Update: June 5, 2016.
Copyright
Publisher
National Center for Biotechnology Information (US), Bethesda (MD)
NLM Citation
Cooper P, Romiti M. Entrez Help. 2006 Jan 20 [Updated 2016 Jun 5]. In: Entrez Help [Internet]. Bethesda (MD): National Center for Biotechnology Information (US); 2005-.