- Exon count:
- 27
| Annotation release |
Status |
Assembly |
Chr
|
Location |
| RS_2023_10 |
current |
GRCh38.p14 (GCF_000001405.40) |
7 |
NC_000007.14 (33129285..33635767) |
| RS_2023_10 |
current |
T2T-CHM13v2.0 (GCF_009914755.1) |
7 |
NC_060931.1 (33268844..33775698) |
| 105.20220307 |
previous assembly |
GRCh37.p13 (GCF_000001405.25) |
7 |
NC_000007.13 (33168897..33675379) |
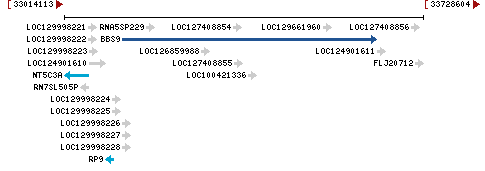
Chromosome 7 - NC_000007.14