Submitting data to GTR in bulk using an excel file with a subset of fields
This document provides detailed information about how to submit data to the GTR describing the clinical tests you offer, via files you prepare in advance. Templates for the files to be uploaded are available from GTR's ftp site. Bulk submission of research tests is not enabled.
There are three major categories of file-based submission that GTR supports or plans to support:
- A file providing details about each test, to be reviewed and submitted via the interactive submission interface (available)
- A file detailing all targets for a specific test, to be uploaded while registering the test, in the methodology tab, on the interactive submission interface (pending)
General Requirements
All submission scenarios require that you first register your laboratory in the GTR using the interactive submission website.
Category 1: Prepare a submission using the GTR test submission template with a subset of fields.
- Familiarize yourself with this submission scenario by reviewing the detailed instructions.
- Download a test submission template file from the GTR ftp site or from your lab's overview page.
- Complete the file according to instructions.
- Use the bulk submission button at the bottom of the overview tab of your lab record to upload your completed file.
- Correct any processing errors, if found. The entire spreadsheet must pass validation; specific error-free tests will not be processed separately.
- Review the test records in the web-based submission interface and supplement information for any recommended and optional fields not supported by the spreadsheet.
- Submit the test records.
Detailed instructions about completing the test submission file and submitting it
- Instructions for completing the test submission file
- Definitions and column values
- Possible error messages
- Steps to upload a completed file
Instructions for completing the file
General features of the test submission file:
This spreadsheet is used to provide a limited set of test-specific data to GTR. Other recommended and optional fields can be completed interactively after successful processing of this form. Registration of a test is not complete until you review, edit and submit each test described on the form. For definitions of minimal, recommended, and optional fields, please go here.
This is a one-time upload of test data that will be used to seed a new record in the GTR. In other words, this file is not meant to be used to submit revisions, updates or to delete existing records. Updates should be managed via the interactive web site or using the full test submission file.
The file does not have to reflect all tests offered by your laboratory. You can prepare and submit multiple test submission files, as long as no test is described in more than one file. You can also enter tests by using the interactive submission site as long as they are distinct from tests submitted by spreadsheet.
Please do not edit rows that start with a # or that have a yellow background.
A red triangle in the upper right of a cell indicates there is a comment. Hover over these cells to see abbreviated instructions on how to complete that cell.
Columns D, E, F, G, L, and N provide pull-down menus. To complete those fields, first click the cell. A gray arrow will appear to the right of the cell. Click on the arrow to see the menu of allowed values and select from it. If you need to select multiple values (e. g. in columns D, F, or G) click on the cell below to add another value. Please note carefully how to manage definitions of a test using multiple rows in the spreadsheet.
If order codes are unique for each test in your lab, and your lab does not have an internal tracking identifier, you can use the order code as your Test tracking ID.
The first column and the first row are frozen as default. However, you may want to unfreeze the first column or freeze another column (e.g. order code) to help keep track of which test you are working on. To do this go to the 'View' tab of the spreadsheet and click the freeze panes icon.
How to enter data:
The form contains 18 columns, or data fields, which you can use to provide data for a test. The first column is critical, because if your submission requires more than one row to describe a test completely, the value in the first column is used to recognize that these data all belong to the same test. In general, each row corresponds to a test target description. A test target is what the test interrogates (e. g. the analyte, chromosomal region/mitochondrion, gene or protein). For simple tests (i.e. one test target), one row corresponds to one test you want to register in GTR. Each set of test targets and linked attributes (e.g. condition the target is assessing, target category, name of what is tested, variants, RefSeq:exons) must be entered in one row so GTR can identify their interrelationship. Thus, a test with multiple test targets will have multiple rows, each with the same value in column A.
Each set of rows with the same Test tracking ID is processed as one test. For single tests that are described using multiple rows, please enter the same Test tracking ID in column A for each applicable row. GTR identifies specific tests via the Test tracking ID, therefore it is crucial that each test be assigned a unique id of your choosing.
Each test target must be linked to at least one indication (condition or phenotype name). Multiple indications for the same test target can be separated by |. Indications placed in the same row as a test target will be connected to that specific target.
If the same indication (condition or phenotype) relates to more than one test target it must be repeated for each appropriate test target row.
Indications (condition or phenotypes) may be entered either as names or identifier (OMIM, CUI, HPO, Orphanet, MeSH, or MONDO). We recommend that you search GTR to find condition/phenotype names recognized by GTR. Use of a recognized condition/phenotype name facilitates discovery of your test through the autocomplete dictionary and disease hierarchies, and enables your test to be linked to clinical, literature and consumer resources.
GTR does not accept registration of send-out tests (i.e. tests that are performed entirely at an outside laboratory/facility as explained here). At least one test component must be performed at the laboratory that is registering the test. If any part of the test is performed at an outside facility, you may enter No for 'Test performance location' on the spreadsheet. When you subsequently review your submission interactively, you should specify which components of the test are performed at an outside facility and provide information in a comment box.
For column/fields that have menu choices, you must select from the choices provided. If multiple choices are applicable, each must be entered in the row(s) below.
If you provide multiple tests for the same disorder, it may help to use the following steps:
- enter one test first,
- use the copy function in your worksheet software to create more rows, and then
- edit only the cells in each row that define what is different about each test, such as the order code.
Definitions and column values
This table describes each value in columns A – Q of the test submission file. The status column (see key) indicates whether the field must be supplied to enable successful upload of the file and the impact on public display.
Two columns (both required) pertain to the spreadsheet submission process: (A) Test tracking ID and (L) Condition/phenotype: choose name or identifier source. All of the remaining columns correspond to fields that are part of overall GTR test submission. The requirement types in overall GTR submission for each column (in parentheses) and field are:
- Minimal: (C) Laboratory test name, (D) Purpose of the test. (E) Test performance location(s), (F) Method Category, (G) Test method, (H) Analytical Validity, (M) Conditions/Phenotypes, (N) Target category, (O) Name of what is tested
- Recommended: (B) Test order code, (I) Citations to support analytical validity, (J) Target population for this test, (K) Citations for target population
- Optional: (P) Variants(s), (Q) RefSeq:relevant exons
Key for Status column:
Private = field will not display to the public
Public = field will be viewable by GTR users
Required = field must be completed for this file to be processed successfully
Recommended = field will display as 'Not provided' if left blank
Optional = field will not display to the public if left blank (i.e. if not applicable for the test)
| Column | Status | Description | Hints |
|---|---|---|---|
| A |
Required |
Test tracking ID. If you have an internal ID, which will help you track your submission, provide it here. You can use your order code if unique for each test or you can come up with your own unique tracking code (e.g. 1,2,3,4,...) | ID to distinguish one test from another on this spreadsheet. This field must be completed for each line. If a single test has multiple lines of data, this field must have the same value for all lines of that test. |
| B | Recommended Public |
Test order code. The order code for this test. | Order code for the test as found on the lab test menu or requisition form that users must use to order this test from this lab. |
| C | Required Public |
Laboratory test name. Your lab name for this test | Test name as found on the lab test menu or requisition form. By default, test name is the record name in GTR and this is how you can easily identify your tests in your submission homepage. Complete this field only for the first line of data for each test. |
| D | Required Public |
Purpose of the test. Select the appropriate purpose or indication of the test. If you need to add multiple test purposes, use a new line for each purpose added. Definitions of choices can be found here. |
In the spreadsheet, you can select from the following list: Diagnosis |
| E | Required Public |
Test performance location(s): Entire test in-house. If the test is performed entirely at the submitting laboratory, select 'Yes'. If any part of the test is performed at an outside location, select 'No'. More information on this field can be found here. |
In the submission interface (not in this spreadsheet) you will be able to be more specific and select from the following options: Entire test performed in-house |
| F | Required Public |
Method Category. Category of the method(s) being used in this test. If multiple methods are used in the test, add an additional line for each method. If your test uses more than one Test method per Method Category, you must select the category again in column F to correspond to the value in column G. To display the complete list of choices, please:
1. Click on the cell - an arrow displays to the right of the cell, then
2. Click on the gray arrow.
|
In the spreadsheet, you can select from the following list: Analyte
Multicolor FISH (M-FISH)/Spectral Karyotyping Chromosome breakage studies Deletion/duplication analysis Enzyme assay FISH-interphase FISH-metaphase Fluorescence in situ hybridization (FISH)
Immunohistochemistry Karyotyping
Linkage analysis Methylation analysis Microsatellite instability testing (MSI) Mutation scanning of select exons Mutation scanning of the entire coding region Protein analysis Protein expression Sequence analysis of select exons Sequence analysis of the entire coding region Sister chromatid exchange Targeted mutation analysis Uniparental disomy study (UPD) |
| G | Required Public |
Test method. Select the test method that corresponds to the entry selected in column 'F' for Method category. Each test must have an entry in both column 'F' and column 'G'. If the test has multiple test methods, add each test method on a new line, making certain the Test method value corresponds to the value in Method category. If the test method used in this test is not listed, please select 'Other' from the list in the spreadsheet and specify the test method during your review of this test record in the interactive submission interface. To display the complete list of choices, please:
1. Click on the cell - an arrow displays to the right of the cell, then
2. Click on the gray arrow.
|
In the spreadsheet, you can select from the following list: Allele-specific primer extension (ASPE)
Alternative splicing detection
Bi-directional Sanger Sequence Analysis C-banding
Chromatin Immunoprecipitation on ChIP Comparative Genomic Hybridization Digital / Virtual karyotyping
Digital microfluidic microspheres
DamID Enzymatic levels Fusion genes microarrays G-banding
Gene expression profiling GeneID Gold nanoparticle probe technology Liquid chromatography-tandem mass spectrometry (LC-MS/MS)
Metabolite levels Methylation-specific PCR
Microarray
Multiplex Ligation-dependent Probe Amplification (MLPA)
Next-Generation sequencing Oligonucleotide Ligation Assay (OLA) Oligonucleotide hybridization-based DNA sequencing PCR-RFLP with Southern hybridization Protein truncation test Pyrosequencing
Q-banding
R-banding
RT-PCR with gel analysis Silver staining
SNP Detection Spectral karyotyping (SKY)
T-banding
Tandem mass spectrometry (MS/MS)
Tiling Arrays Trinucleotide repeat by PCR or Southern Blot Uni-directional Sanger sequencing Other |
| H | Required Public |
Analytical Validity. Required once per test. Provide quantitative information about analytical sensitivity, analytical specificity, accuracy, and/or precision of this test. Example: 'The OtoChip is greater than 99% sensitive for detecting substitution variants in the sequence analyzed. In addition, this assay is 95% sensitive for detecting small insertions or deletions (InDels) (100% for 1-2 bp indels, 92.3% for 3-5 bp indels and 87.5% for >10 bp indels)'. |
The Analytical validity field needs a statement that includes quantitative information for analytical sensitivity, analytical specificity, accuracy, and/or precision of this test. General statements about methodologies that do not contain specific information about the test do not satisfy the specifications for this field. Nor do statements such as "N/A", "not applicable", "Yes" or "unknown". |
| I | Recommended Public |
Citations to support analytical validity. PubMed identifiers preferred; otherwise, full citations or URLs. If multiple, separate by '|'. | Provide PubMed ID when available. This will allow GTR to provide a link for users to have easy access to the citation. If no publications are available, provide non-proprietary internal lab data in analytical validity statement, if available. You can also provide a link to your lab's webpage with information about analytical validity for this test. |
| J |
Recommended
Public
|
Target population for this test. Provide information on which population segment(s) the test is appropriate for and why. Complete this field once for each test. | Example: Individuals with clinical symptoms consistent with Noonan syndrome. |
| K |
Recommended
Public
|
Citations for target population. PubMed identifiers preferred; otherwise, full citations or URLs. If multiple, separate by '|'. | Provide PubMed ID when available. This will allow GTR to provide a link for users to have easy access to the citation. You can also provide a link to your lab's webpage with information about the target population for this test. |
| L |
Required
Private
|
Condition/phenotype: choose name or identifier source Select whether the condition/phenotype will be provided by condition name or identifier (OMIM, CUI, HPO, Orphanet, MeSH, or MONDO). To display the list of choices, please:
1. Click on the cell - an arrow displays to the right of the cell, then
2. Click on the gray arrow.
|
In the spreadsheet, you can select from the following list:
condition name |
| M | Required Public |
Conditions/Phenotypes If you selected MIM number in previous column (L), enter the valid MIM condition number (please do not enter MIM number for gene records). If you enter a MIM number for the condition being tested, GTR will use this to match your submission to disorders already represented in GTR's database. Similarly, if you selected one of the other identifier sources in Column L, enter the valid identifier here. If you selected condition name, write the preferred name of the condition/phenotype. If multiple conditions/phenotypes exist for a single target (columns N, O and P), enter values separated by a vertical bar (|). If same condition/phenotype exists for multiple targets, the condition/phenotype must be repeated on each target line. Preferred names can be found by searching GTR. |
Valid MIM condition number (please do not enter MIM number for gene records) or identifier corresponding to the identifier source. If you enter a MIM number for the condition being tested, GTR will use this to match your submission to disorders already represented in GTR's database. The list of names GTR uses per MIM number is provided from the ClinVar's FTP site. You can search the GTR to find the condition name in the database. Linking to an existing condition name ensures that this test will be connected to available resources such as clinical and molecular resources. Learn how to use MedGen to select condition and phenotype terms. |
| N | Required Public |
Target category. Select the category describing the test target measures in the test. If multiple targets are being tested, add each target on a separate line. To display the complete list of choices:
1. Click on the cell - an arrow displays to the right of the cell, then
2. Click on the gray arrow.
|
In the spreadsheet, you can select from the following list: analyte Note that gene is used to represent molecular assays that detect variations in a gene or set of genes. |
| O | Required Public |
Name of what is tested. If gene in column N, enter HGNC official gene symbol. If chromosomal region/mitochondrion in N, enter region. If protein, please use Swiss-Prot protein name. If an analyte, enter its name. If multiple targets are being tested, each target must be stored on a separate line. |
Examples: Analyte: 'Branched-chain amino acids' or the enzyme 'Branched-chain ketoacid dehydrogenase' Chromosomal region/mitochondrion: '11q15' or 'm.7446-m.7514' Gene: 'CDKN2A' or 'CFTR' Protein: 'Hemoglobin subunit beta' or 'Adenomatous polyposis coli protein' |
| P | Optional Public |
Variants(s). Describe variation using fully qualified HGVS expression,e.g. NM_000258.2:c.427G>A. If multiple variants for Conditions/phenotypes named in M, separate with '|'. |
Fill this column only if gene is provided in columns N. Example: for a test that interrogates the 1477_1478delCA and 1477C>T of the CFTR gene for Cystic fibrosis, you would enter: Column L: MIM number
Column M: 219700
Column N: gene
Column O: CFTR
Column P: NM_000492.3:c.1477_1478delCA|NM_000492.3:c.1477C>T
|
| Q | Optional Public |
RefSeq:relevant exons. Provide details about the exons analyzed and specify the associated reference sequences. Format by first specifying reference sequence, include assession number and version, then using ':', and then listing exons. If multiple reference sequences per gene, separate these by '|'. |
Valid only if gene is provided in column N. The RefSeq defining the exon numbering system must be supplied in this column. Example: NM_4433221.1:1-5(all)|NM_123456.1:1,3,4-5 including 5 bp in intron-exon boundaries. If a continuous range is tested: provide the exon numbers as a range, e.g. 2-5. If the range includes all exons, add (All), e.g. 1-5 (All) If a subset is tested: enumerate the exons tested. Ranges are allowed, e.g.3,5,9-12 NOTE: If gene name is provided, the chromosomal location (RefSeq) will be automatically provided and does not need to be entered UNLESS you are providing exons. |
Possible error messages
The following is a list of errors that may arise when your file is processed. These are the validation points that are checked and we encourage you to be aware of them so that you can avoid them when completing your file. When a file is uploaded with data that is not accepted or with missing data, you will receive an email listing the errors encountered in the file so you can make the appropriate edits. The possible error messages are:
-
Cell with missing values (required fields only, for a list of required fields, please see the table 'Definitions and column values').
-
Cell contains multiple values whereas only a single value is allowed.
-
The number of values between certain cells is required to match. Some cells are interdependent; for example, each test target needs to be linked to at least one indication.
-
The condition/phenotype name is not recognized by GTR. Please check the syntax or spelling of the terms you enter. You can find recognized terms by searching GTR.
-
The MIM number is not recognized as a condition record. Please make sure that all MIM numbers correspond to a condition/phenotype record in OMIM (i.e. MIM numbers for gene records are not accepted as condition identifiers).
-
The entry for 'Name of what is tested' does not match a gene symbol in the GTR database, which uses symbols from the HUGO Gene Nomenclature Committee (HGNC). Please review entries for misspellings and/or search GTR for recognized gene symbols.
Processing also checks for order codes that match tests already registered by your lab in GTR and if found, will generate a warning message. This is not considered to be an error. Laboratories may use the same order code for different tests, so this check is meant to alert you to the possibility of duplication or a typographical error in entering a code. This message is information for your consideration and will not halt processing of your file.
Follow these steps to upload a completed file
Step 1. Download the file gtr_test_submission_template.xlsx
Step 2. Complete the file
Step 3. Go to the GTR submission user interface and Login. You may used your NIH login or third-party login credentials.
Step 4. Click on your laboratory name or the link 'Review lab submission' to go to your lab record
Step 5. Go to the Overview tab of your lab record
Step 6. Scroll to the bottom of the page (or use the End button on your keyboard)
Step 7. Upload the completed file.
- Browse your computer
- Select the file
- Click the 'Upload your bulk submission file' button
Step 8. Notification of submissions - you will receive a notification via email within a few minutes of uploading your file:
- If your spreadsheet is successfully uploaded, you will receive an email entitled 'Your tests submitted to GTR in bulk via spreadsheet are ready for review'.
- If your speadsheet is successfully uploaded but has tests with order codes that match tests you already submitted to GTR, you will receive an email entitled 'Your tests submitted to GTR in bulk via spreadsheet are ready for review' with a section where 'warnings' are discussed. 'Warnings' are a list of tests that have the same order codes as tests already in the GTR. You can review these tests in the submission user interface, in case they were submitted in error as duplicates or the order codes need to be edited. Distinct tests can have the same order code, so you are not required to make edits. This validation step is performed to help you detect possible duplications in your submissions.
- If your spreadsheet contains errors and cannot be processed, you will receive error messages in the user interface and an email entitled 'Your tests submitted to GTR in bulk via spreadsheet contain processing errors which require your attention'. You will need to correct these errors and resubmit your spreadsheet. A list of errors and how to address them can be found above.
Step 9. The file you uploaded will display in the 'History of bulk test uploads' and is accessible to you. You can download that file if you need to address errors or edit data. You will also notice the date and time of upload. Please consider versioning your template if you need to make corrections so that you can later distinguish which version you uploaded to GTR.
Step 10. Refresh the page.
- Once the spreadsheet has been successfully processed, the tests uploaded will display in your GTR submission homepage.
- There is a status box on the table that tells you the file's processing status: 'success', 'warnings' or 'error'.
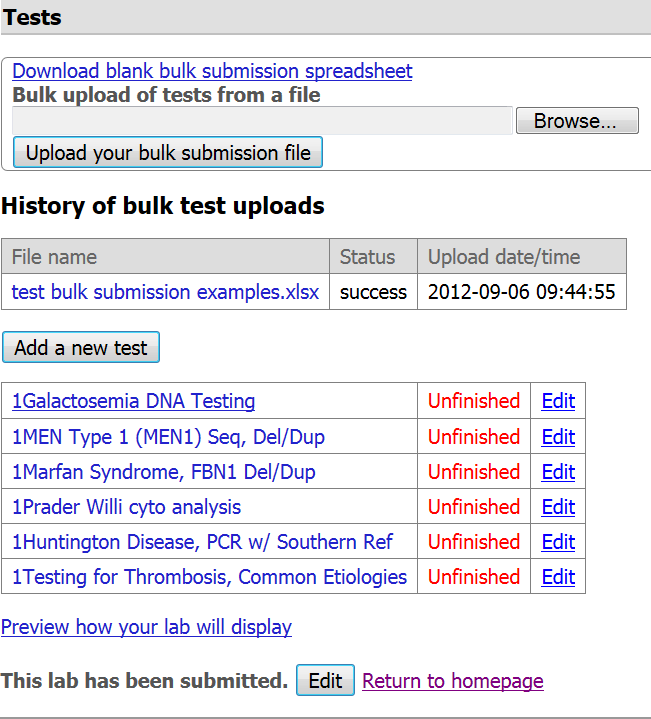
Step 11. You need to click on each test name (or the "Edit" button next to the test name) to review the data about each test record. We encourage you to complete information on recommended and optional fields not supported by the test submission file while reviewing your data in the interactive submission interface. Fields that may need special attention:
- If you entered 'Other' for Test methodology (column G), please enter the value in the Methodology tab
- If you entered 'No' for Test performance location: Entire test in-house (column E), please make your specific selections and provide information in the comment box in the Performance characteristics tab
Step 12. Preview how your test will display to the public before submitting it by clicking the link 'Preview how your test will display'. This enables you to return to the test record to edit information before your test is displayed publicly. For more information on how to review your test records please go here.
Step 13. Submit your test record by clicking the 'Submit' button. Assuming normal processing, your tests will be publicly viewable within 24-48 hours. Check out your tests on the public site and evaluate the test menu in your laboratory page. You can edit the data as necessary in your submission user interface.
A file providing the details about each target your test detects (pending)
This feature will enable explicit descriptions of the genes and other targets assayed by your test. Currently GTR supports submission of these data by using the minimal or full test submission templates or by entering targets individually using interactive forms. The target file will enable upload of a tab-delimited file providing all the details for a specific test within the test Methodology tab in the interactive submission interface.