#include <corelib/ncbiobj.hpp>#include <corelib/ncbistd.hpp>#include <corelib/ncbi_limits.hpp>#include <set>#include <vector>#include <algorithm>#include <math.h>#include <objmgr/seq_vector_ci.hpp>#include <util/range.hpp> Include dependency graph for gnomon_model.hpp:
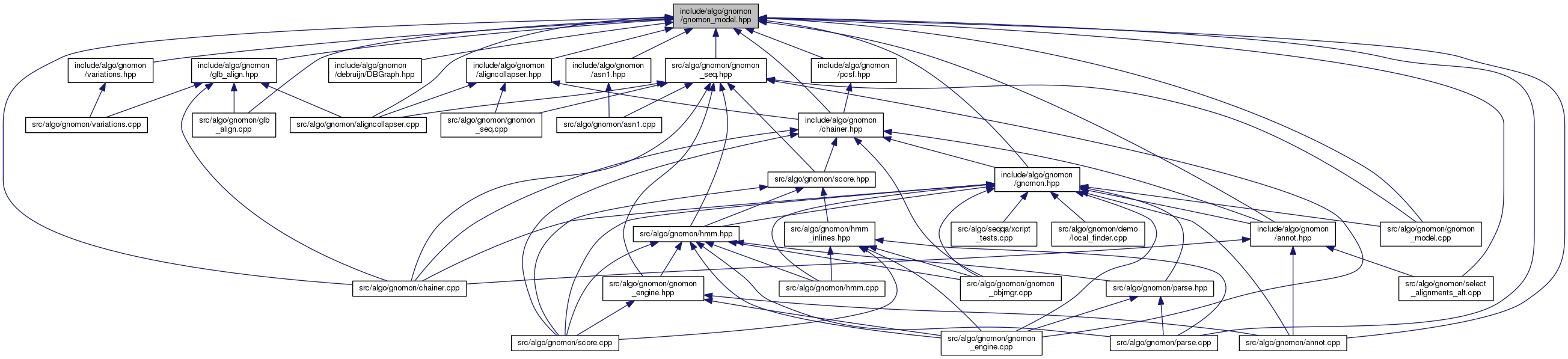
Include dependency graph for gnomon_model.hpp: This graph shows which files directly or indirectly include this file:
This graph shows which files directly or indirectly include this file:Go to the source code of this file.
Go to the SVN repository for this file.
Classes | |
| class | CSupportInfo |
| class | CInDelInfo |
| struct | CInDelInfo::SSource |
| class | CRangeMapper |
| class | CModelExon |
| class | CCDSInfo |
| struct | CCDSInfo::SPStop |
| class | CGeneModel |
| class | CAlignMap |
| struct | CAlignMap::SMapRangeEdge |
| class | CAlignMap::SMapRange |
| class | CAlignModel |
| struct | setcontig |
| struct | getcontig |
| class | CModelCluster< Model > |
| class | CModelClusterSet< Cluster > |
| class | EResidue |
Typedefs | |
| typedef objects::CSeqVectorTypes::TResidue | TResidue |
| typedef vector< TResidue > | CResidueVec |
| typedef vector< int > | TIVec |
| typedef vector< double > | TDVec |
| typedef set< CSupportInfo > | CSupportInfoSet |
| typedef vector< CInDelInfo > | TInDels |
| typedef CModelCluster< CGeneModel > | TGeneModelCluster |
| typedef CModelCluster< CAlignModel > | TAlignModelCluster |
| typedef list< CGeneModel > | TGeneModelList |
| typedef list< CAlignModel > | TAlignModelList |
| typedef CModelClusterSet< TGeneModelCluster > | TGeneModelClusterSet |
| typedef CModelClusterSet< TAlignModelCluster > | TAlignModelClusterSet |
Enumerations | |
| enum | EStrand { ePlus , eMinus } |
| enum | EResidueNames { enA , enC , enG , enT , enN } |
Variables | |
| const EResidue | k_toMinus [5] |
| const char *const | k_aa_table |
Typedef Documentation
◆ CResidueVec
| typedef vector<TResidue> CResidueVec |
Definition at line 69 of file gnomon_model.hpp.
◆ CSupportInfoSet
| typedef set<CSupportInfo> CSupportInfoSet |
Definition at line 96 of file gnomon_model.hpp.
◆ TAlignModelCluster
| typedef CModelCluster<CAlignModel> TAlignModelCluster |
Definition at line 814 of file gnomon_model.hpp.
◆ TAlignModelClusterSet
Definition at line 839 of file gnomon_model.hpp.
◆ TAlignModelList
| typedef list<CAlignModel> TAlignModelList |
Definition at line 817 of file gnomon_model.hpp.
◆ TDVec
| typedef vector<double> TDVec |
Definition at line 71 of file gnomon_model.hpp.
◆ TGeneModelCluster
| typedef CModelCluster<CGeneModel> TGeneModelCluster |
Definition at line 813 of file gnomon_model.hpp.
◆ TGeneModelClusterSet
Definition at line 838 of file gnomon_model.hpp.
◆ TGeneModelList
| typedef list<CGeneModel> TGeneModelList |
Definition at line 816 of file gnomon_model.hpp.
◆ TInDels
| typedef vector<CInDelInfo> TInDels |
Definition at line 169 of file gnomon_model.hpp.
◆ TIVec
Definition at line 70 of file gnomon_model.hpp.
◆ TResidue
| typedef objects::CSeqVectorTypes::TResidue TResidue |
Definition at line 67 of file gnomon_model.hpp.
Enumeration Type Documentation
◆ EResidueNames
| enum EResidueNames |
| Enumerator | |
|---|---|
| enA | |
| enC | |
| enG | |
| enT | |
| enN | |
Definition at line 842 of file gnomon_model.hpp.
◆ EStrand
| enum EStrand |
| Enumerator | |
|---|---|
| ePlus | |
| eMinus | |
Definition at line 64 of file gnomon_model.hpp.
Function Documentation
◆ BadScore()
|
inline |
Definition at line 62 of file gnomon_model.hpp.
References max().
Referenced by CGnomonEngine::AcceptorScore(), AddProbabilities(), CIntergenic::BranchScore(), CInternalExon::BranchScore(), CSingleExon::BranchScore(), CLastExon::BranchScore(), CFirstExon::BranchScore(), CIntron::BranchScore(), CalcStateScores(), CGeneModel::CdsInvariant(), CCDSInfo::Clear(), CCDSInfo::Clip(), CExon::ClosingLengthScore(), CLorentz::ClosingScore(), CCDSInfo::CombineWith(), CParse::CParse(), CAnnotationASN1::CImplementationData::create_internal_feature(), CChainer::CChainerImpl::CreateChainsForPartialProteins(), CCDSInfo::Cut(), CGnomonEngine::DonorScore(), CChainer::CChainerImpl::Duplicate5pendsAndShortCDSes(), CChainer::CChainerImpl::DuplicateUTRs(), CChainer::CChainerImpl::FilterOutBadScoreChainsHavingBetterCompatibles(), CChainer::CChainerImpl::FindGeneSeeds(), CChainer::FindSelenoproteinsClipProteinsToStartStop(), CCodingPropensity::GetScore(), CGnomonEngine::GetScore(), CChainer::CChainerImpl::GoodCDNAScore(), CSeqScores::Init(), CExon::InitialLengthScore(), CChainMembers::InsertMember(), CCDSInfo::Invariant(), CSeqScores::isConsensusIntron(), CIntron::LengthScore(), CChainer::CChainerImpl::MakeChains(), MemberIsCoding(), SingleExon_Noncoding::model_predicate(), LowSupport_Noncoding::model_predicate(), operator>>(), CGnomonEngine::PointCodingScore(), CGnomonAnnotator::Predict(), SGFFrec::print(), CGnomonAnnotatorArgUtil::ReadArgs(), readGFF3(), CGnomonEngine::Run(), s_EvaluateNewScore(), s_ForwardStep(), s_MakeStep(), s_TooFar(), CWAM_Donor< order >::Score(), CWAM_Acceptor< order >::Score(), CWMM_Start::Score(), CWAM_Stop::Score(), CMC_NonCodingRegion< order >::Score(), CMC3_CodingRegion< order >::Score(), CChainer::CChainerImpl::ScoreCdnas(), CChainer::ScoreCDSes_FilterOutPoorAlignments(), CGnomonEngine::SelectBestReadingFrame(), CIntronParameters::SetSeqLen(), CIntergenicParameters::SetSeqLen(), CLorentz::Through(), CExon::ThroughLengthScore(), CMarkovChain< 0 >::toScore(), CGnomonAnnotator::TryToEliminateAlignmentsFromTail(), CGnomonAnnotator::TryToEliminateOneAlignment(), and CGnomonAnnotator::TryWithoutObviouslyBadAlignments().
◆ Complement() [1/2]
Definition at line 883 of file gnomon_model.hpp.
References k_toMinus.
◆ Complement() [2/2]
Definition at line 855 of file gnomon_model.hpp.
Referenced by ReverseComplement(), and DeBruijn::CDBGraphDigger::SContig::SContig().
◆ Enclosed()
|
inline |
Definition at line 77 of file gnomon_model.hpp.
References Include().
◆ GetAlignParts()
Definition at line 897 of file gnomon_model.hpp.
References CGeneModel::eRemoveExons, CGeneModel::Exons(), i, ncbi::grid::netcache::search::fields::size, and CGeneModel::Status().
Referenced by CAlignCollapser::AddAlignment(), CChainer::CChainerImpl::CutParts(), and CAlignCollapser::FilterAlignments().
◆ Include() [1/2]
|
inline |
Definition at line 75 of file gnomon_model.hpp.
References CRange_Base::GetFrom(), and CRange_Base::GetTo().
Referenced by AddSupport(), BelongToExon(), CChain::CalculateDropLimits(), CChain::CalculateSupportAndWeightFromMembers(), CChainer::CChainerImpl::CanIncludeJinI(), CGeneModel::CdsInvariant(), CChain::CheckSecondaryCapPolyAEnds(), CMultAlign::CheckWord(), CAlignCollapser::CleanSelfTranscript(), CChain::ClipChain(), CChain::ClipLowCoverageUTR(), CChain::ClipToCap(), CChain::ClipToPolyA(), CGeneModel::CombineCdsInfo(), CChainer::CChainerImpl::CombineCompatibleChains(), CCDSInfo::CombineWith(), CSeqScores::ConstructSequenceAndMaps(), CAnnotationASN1::CImplementationData::create_cdregion_feature(), CChainer::CChainerImpl::CreateChainsForPartialProteins(), CCDSInfo::Cut(), Enclosed(), CAlignCollapser::FilterAlignments(), CChainer::CChainerImpl::FindContainedAlignments(), CChainer::CChainerImpl::FindOptimalChainForProtein(), FindStartsStops(), CGeneModel::FShiftedLen(), CAlignMap::FShiftedLen(), CGnomonEngine::GetScore(), CGene::HarborsRange(), CSeqScores::Init(), CCDSInfo::Invariant(), CGeneModel::IsSubAlignOf(), CChainer::CChainerImpl::LRCanChainItoJ(), CChainer::CChainerImpl::LRIinit(), CChain::MainPeaks(), CChainer::CChainerImpl::MakeChains(), SChainMember::MarkUnwantedCopiesForChain(), CGnomonAnnotator::Predict(), CModelCompare::RangeNestedInIntron(), CChain::RemoveFshiftsFromUTRs(), CChainer::CChainerImpl::ReplicatePStops(), CChain::RestoreReasonableConfirmedStart(), CChainer::CChainerImpl::RightLeft(), CMultAlign::SeqCountsBetweenTwoStrongWords(), CChain::SetConfirmedEnds(), CChain::SetConfirmedStartStopForCompleteProteins(), CChain::SetConsistentCoverage(), CAlignMap::ShrinkToRealPoints(), CAlignMap::ShrinkToRealPointsOnEdited(), CChainer::CChainerImpl::TrimAlignmentsIncludedInDifferentGenes(), and CGnomonAnnotator::TryWithoutObviouslyBadAlignments().
◆ Include() [2/2]
|
inline |
Definition at line 76 of file gnomon_model.hpp.
References r().
◆ IsStartCodon()
Definition at line 108 of file gnomon_seq.cpp.
References res_traits< Res >::_codons(), res_traits< Res >::_rev_codons(), and ePlus.
Referenced by FindStartsStops(), CChain::SetConfirmedEnds(), and ProjectCDS::transform_align().
◆ IsStopCodon()
Definition at line 124 of file gnomon_seq.cpp.
Referenced by CGnomonEngine::GetScore(), Partial5pCodonIsStop(), CChain::SetConfirmedEnds(), and ProjectCDS::transform_align().
◆ MapAlignsToOrigContig()
| void MapAlignsToOrigContig | ( | TAlignModelList & | aligns, |
| const TInDels & | corrections, | ||
| int | contig_size | ||
| ) |
◆ operator<<() [1/3]
| CNcbiOstream& operator<< | ( | CNcbiOstream & | s, |
| const CAlignModel & | a | ||
| ) |
Definition at line 2566 of file gnomon_model.cpp.
References a, eGFF3FileFormat, model_file_format_state, and printGFF3().
◆ operator<<() [2/3]
| CNcbiOstream& operator<< | ( | CNcbiOstream & | s, |
| const CGeneModel & | a | ||
| ) |
Definition at line 2561 of file gnomon_model.cpp.
References a, and operator<<().
◆ operator<<() [3/3]
| CNcbiOstream& operator<< | ( | CNcbiOstream & | s, |
| const setcontig & | c | ||
| ) |
Definition at line 1605 of file gnomon_model.cpp.
References contig_stream_state, and setcontig::m_contig.
Referenced by operator<<().
◆ operator>>() [1/2]
| CNcbiIstream& operator>> | ( | CNcbiIstream & | s, |
| CAlignModel & | a | ||
| ) |
Definition at line 2577 of file gnomon_model.cpp.
References eGFF3FileFormat, model_file_format_state, and readGFF3().
◆ operator>>() [2/2]
| CNcbiIstream& operator>> | ( | CNcbiIstream & | s, |
| const getcontig & | c | ||
| ) |
Definition at line 1609 of file gnomon_model.cpp.
References contig_stream_state, and getcontig::m_contig.
◆ OtherStrand()
Definition at line 65 of file gnomon_model.hpp.
Referenced by isGoodIntron().
◆ Precede()
|
inline |
Definition at line 74 of file gnomon_model.hpp.
References CRange_Base::GetTo(), and r().
Referenced by CModelCompare::CanBeConnectedIntoOne(), CCDSInfo::Clear5PrimeCdsLimit(), CCDSInfo::CombineWith(), CCDSInfo::Invariant(), CCDSInfo::MapFromEditedToOrig(), CCDSInfo::MapFromOrigToEdited(), Precedence::operator()(), RangeOrder::operator()(), CGeneModel::operator<(), CModelCluster< Model >::operator<(), CModelExon::operator<(), CGnomonAnnotator::Predict(), CChain::RestoreReasonableConfirmedStart(), CCDSInfo::SetStop(), CCDSInfo::Strand(), and CChainer::CChainerImpl::TrimAlignmentsIncludedInDifferentGenes().
◆ ReverseComplement()
| void ReverseComplement | ( | const BidirectionalIterator & | first, |
| const BidirectionalIterator & | last | ||
| ) |
Definition at line 889 of file gnomon_model.hpp.
References Complement(), first(), i, and last().
Referenced by CDiscrepancyVisitorImpl< _Name >::Autofix(), BOOST_AUTO_TEST_CASE(), CAlignCollapser::CleanSelfTranscript(), CAlignCollapser::FillGapsInAlignmentAndAddToGenomicGaps(), CGeneModel::GetCdsDnaSequence(), CCorrectRNAStrandDlg::GetCommand(), CMixedStrands::GetCommand(), CParse::GetGenes(), GetReverseComplimentSequenceCommand(), CGnomonAnnotator_Base::MapOneModelToEditedContig(), CGnomonAnnotator_Base::MapOneModelToOrigContig(), CRevCompSequencesDlg::RevCompBioSeq(), DeBruijn::CDBGraphDigger::SContig::ReverseComplement(), CGeneModel::ReverseComplementModel(), and CChain::ValidPolyA().
◆ USING_SCOPE()
| USING_SCOPE | ( | objects | ) |
Variable Documentation
◆ k_aa_table
Definition at line 41 of file gnomon_seq.cpp.
◆ k_toMinus
Definition at line 40 of file gnomon_seq.cpp.
Referenced by Complement(), CSeqScores::Init(), and ReverseComplement().